Research Article
Volume 1 Issue 1 - 2017
Abiotic Stress Tolerance in Crop Plants: With Respect to NAC Transcription Factor
Department of Ag. Biotechnology, College of Agriculture, S.V.P. University of Agriculture & Technology, Meerut 250 110 India
*Corresponding Author: Ashu Singh, Department of Ag. Biotechnology, College of Agriculture, S.V.P. University of Agriculture &
Technology, Meerut 250 110 India.
Received: December 07, 2016; Published: June 19, 2017
Abstract
Natural factors such as salinity, drought and high temperatures are the main cause of low productivity of crop plants worldwide. Plants respond to such extremities through various mechanisms. Thus an insight into the various complex mechanisms can broaden our understanding and can be a remedy to the obscurity of abiotic stress tolerance in plants. Many workers all over the world have tried to unveil the gene networks responsible for abiotic stress tolerance. After the discovery of transcription factors, it was clear that abiotic stress tolerance is modulated by numerous genes which get up regulated during stress conditions. TFs are therefore good candidates for molecular genetics to improve crop tolerance to abiotic stress because of their role as major regulators of clusters of genes. Thus these collective efforts and results of collaborative studies will positively contribute to sustainable food production in the world and will help to prevent global-scale environmental damage that results from abiotic stress. In this review NAC transcription factor genes utilized for development of transgenic abiotic stress tolerant plants and will be discussed
Keywords: Transcription factors; Abiotic stress; ROS; NAC; DREB; ARF; Protein
Abiotic Stresses: Abiotic stresses such as drought, temperature, and salinity are location specific, displaying inner variation in occurrence, intensity and duration, and generally cause reduced crop productivity. There is a necessary concern for food security in developing countries including India for the following reasons: (i) The food demand is increasing for the ever growing population; (ii) declining productivity in past high productivity regions, often described as 'Green Revolution' fatigue; and (iii) increasing susceptibility of agriculture to potential climate change. Climatic change is becoming a large concern towards humanity in coming decades. Agriculture also produces significant effects on climate change as possible contributor of greenhouse gases to the atmosphere and as an industry that is highly sensitive to climatic changes. Climate is significant in the distribution, productivity and security of food.
There should be a realization that climate is both a resource to be managed wisely and a hazard to be dealt with. Thus a portfolio of assets to prepare for climate change is needed which is a major cause abiotic aberration. Of all the parameters, abiotic environmental stresses contribute most significantly to the reduction in potential yield (Flowers & Yeo, 1995). To deal with scarce good quality water resources, agronomic research activity has focused on the optimization of water use for irrigation and on the use of alternative sources of waters. Breeders take a relatively traditional approach. They grow and cross varieties then evaluate how the progenies vary in their ability to deal with stresses.
The best-adapted plants will then be selected for growing in fields exposed to stresses. Biotechnologists, meanwhile, have taken advantage of recent advances in functional genomics and biotechnology to genetically engineer crops which can give better yield than the unmodified ones in adverse conditions (Katerji., et al. 2003; Katerji., et al. 2004; Manavalan., et al. 2009; Umezawa., et al. 2006). The genetic basis of stress tolerance is complex and not well understood although many attempts are being made (Yadav., et al. 2013). The ability of plants to adapt and/or acclimate to different environments is directly or indirectly related with the plasticity and resilience of photosynthesis, in combination with other processes, determining plant growth and development, namely reproduction (Chaves., et al. 2011). The most striking feature of plant adaptation to abiotic stresses is the activation of multiple responses involving complex gene interactions and crosstalk with many molecular pathways (Basu 2012; Umezawa., et al. 2006). Abiotic stresses elicit complex cellular responses that have been elucidated by progresses made in exploring and understanding plant abiotic responses at the whole-plant, physiological, biochemical, cellular and molecular levels (Grover 2001). At the molecular level this involves induction of stress-responsive and stress-tolerance genes (Matsui., et al. 2008), often mediated by the phytohormone abscisic acid (ABA). ABA is referred to as the plant stress hormone because, in addition to its role in development, it plays a key role in responses to abiotic stress factors by regulating stomatal closure to optimize transpiration, and by triggering the activation of many stress-related genes (Cutler., et al. 2008) (Finkelstein., et al. 2002).
Several genes are activated in response to abiotic stresses at the transcriptional level, and their products are contemplated to provide stress tolerance by the production of vital metabolic proteins and also in regulating the downstream genes (Kavar., et al. 2007). Extensive transcriptome analyses have divulged that these gene products can largely be classified into two groups (Bohnert., et al. 2001; Seki., et al. 2002; Fowler and Thomashow 2002), First group [Figure 1] comprises of genes that encode for proteins that defend the cells from the effects of water-deficit. These genes mainly include those that regulate the accumulation of compatible solutes (enzymes for osmolyte biosynthesis like proline, betaine, sugars, etc.); passive and active transport systems across membranes (water channel proteins and membrane transporters); and protection and stabilization of cell structures from damage by ROS (the detoxification enzymes such as glutathione S-transferase, catalase, superoxide dismutase , ascorbate peroxidase, etc.); fatty acid metabolism enzymes, proteinase inhibitors, ferritin and lipid-transfer proteins; and other proteins for the protection of macromolecules (LEA protein, osmotin, chaperons, etc.). Another group of genes stimulated by abiotic stresses includes regulatory proteins that further regulate the stress signal transduction and alter gene expression and hence possibly function in stress response. They comprise several transcription factors (TFs) emphasizing the role of various transcriptional regulatory mechanisms in the stress signal transduction pathways; protein kinases (MAP kinase, CDP kinase, receptor protein kinase, etc.); protein phosphatases and proteinases implicated in the regulation of stress signaling and gene expression (Seki., et al. 2003; Shinozaki and Yamaguchi- Shinozaki, 2007).
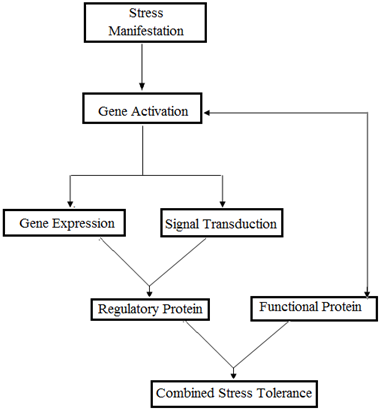
Figure 1: Diagrammatic representation of stress tolerance by two groups of Proteins and their combined action.
As we know that classical tools of genetics and plant breeding have already confirmed that abiotic stress tolerance in plants is multigenic and quantitative. Hence, it is difficult to manipulate abiotic-stress-related pathways using modern molecular genetics techniques. Introduction of a single gene, encoding functional proteins like LEA proteins, antifreeze proteins, and molecular chaperons, would confer some degree of tolerance but do not give sustained tolerance to most of the abiotic stresses. However, during the evolutionary process, plants have devised some intense molecular mechanisms in order to survive under the extreme environmental conditions. In this way, another category of genes namely genes encoding regulatory proteins have emerged. Such genes play important roles in survival of plants under stress situation by acting as a switch of sets of downstream stress-responsive genes.
Recent advances in biotechnology have dramatically changed our capabilities for gene discovery and functional genomics. For the first time, we can now obtain a holistic “snapshot” of a cell with transcript, protein and metabolite profiling. In plant genomes approximately 7% of the coding sequences are assigned to transcription factors (TFs) (Udvardi., et al. 2007), and many of these are immediate-early abiotic stress-responsive genes (Kilian., et al. 2012). [Figure 2] represents the key factors responsible for stress tolerance. The Abiotic stress (Cold, drought, salinity and heat) induces subsequent signal transduction through either ABA-dependent or ABA-independent pathways, leading to the activation of various physiological and metabolic responses leading to activation of NAC Genes thus subsequent production of NAC mRNA and finally after translation production of NAC TF protein. These TFs interact with the conserved “stress recognition sequences” present as cis-acting elements upstream the promoter of the concerned gene, thereby activating the gene responsible for stress tolerance. Manipulation of these genes should be the major target of attempts to produce plants with enhanced multiple stress tolerance. Furthermore, the potential NAC genes which confer multiple abiotic stress tolerance in model plant species must be tested in crop plants and greater emphasis should be placed on the field evaluation of the transgenic crops harboring these genes, especially focusing on the productive success.
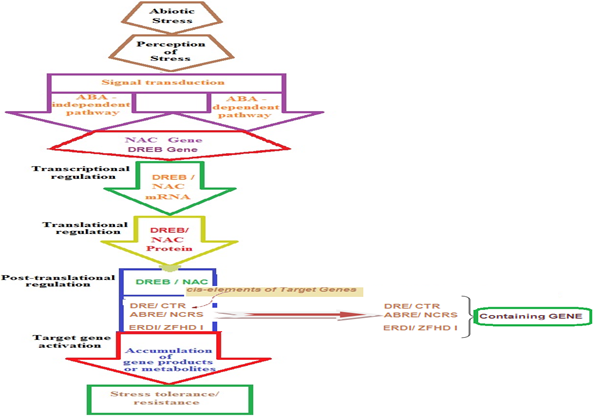
Figure 2: Schematic diagram of NAC TF as key components in
transcriptional regulatory networks during abiotic stress.
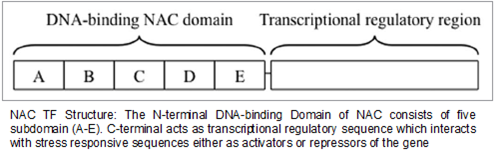
Structure and Function of TF with reference to NAC: The expression of inducible genes in crops is controlled by specific proteins known as transcription factors at the transcriptional level. A typical transcription factor usually contains a DNA-binding domain, a transcription regulation domain, a dimerization site and a nuclear localization domain. These functional domains define the characteristic, localization and regulatory role of a transcription factor. Transcription factors recognize and bind to specific cis-acting elements or interact with other proteins, and then activate or repress the transcription of target genes by their functional domains. In recent years, elucidation on the structure and function of transcription factors has become an important subject in plant molecular biology. Thus all TFs have a DNA-binding domain containing a short peptide region, called the DNA-binding motif. The NAC domain (for Petunia hybrida (Petunia) NAM and for Arabidopsis ATAF1, ATAF2, and CUC2) is an N-terminal module of ~160 amino acids, the NAC domain is accompanied by diverse C-terminal transcriptional activation domains. The NAC domain has been shown to be a DNA-binding domain (DBD) and a dimerization domain (www.ebi.ac.uk/interpro/entry/IPR003441). Regulation of transcription often works through binding of regulatory proteins to the cis-acting elements located upstream of functional genes. The C-terminal parts of NAC proteins are highly diverse and do not contain any known protein domains (Ooka., et al. 2003).
However, sequence analysis does reveal five Arabidopsis NAC genes that appear to encode two NAC domains in tandem (At1g60280, At1g60300, At1g60340, At1g60350 and At1g60380) as well as sequences predicted to contain transmembrane regions or nuclear import and/or export signals. Elucidation of the molecular functions of NAC proteins began with the report of two NAC proteins being able to activate a cauliflower mosaic virus (CaMV) 35S promoter construct in yeast (Xie., et al. 2000). Since then, three NAC proteins from Arabidopsis (NAC1, AtNAM and ANAC019) and several from Brassica napus have been shown to bind the CaMV 35S promoter (Xie., et al. 2000; Hegedus., et al. 2003; Ernst., et al. 2004; Duval., et al. 2002). Recently, three ANAC proteins were shown to bind a fragment of the ERD1 (EARLY RESPONSIVE TO DEHYDRATION STRESS 1) promoter. Substitution analysis identified the sequence CACG as the core DNA motif recognized by the ANAC proteins in the promoter fragment (Tran., et al. 2004). The DNA-binding ability has been assigned to the NAC domain (Hegedus., et al. 2003; Ernst., et al. 2004; Duval., et al. 2002), but the mode of DNA recognition by the NAC domain is still unknown.
Bioengineering for Improving Stress Tolerance: Engineering plants for overexpression of genes encoding for transcriptional factors are identified as an effective method for generating abiotic stress-tolerant plants. Gene expression studies using constitutive promoters provide very little biological information compared with the use of inducible promoters or cell type-specific promoters. The choice of promoters can significantly affect the results from a transgenic manipulation; basic molecular biology research has established that expression of a given gene is governed by the promoter sequence present mostly at 5’ end of gene. The promoter sequences determine the strength of the expression (i.e. the extent to which the gene will be expressed) as well as the preciseness of the quadrants to which the gene will be expressed (temporal and/or spatial). For instance, regulation of any gene for example Heat shock (HS) gene is mediated by a core DNA sequence called heat shock element (HSE), located in the promoter region of the hs gene which upon stimulation by stress condition transcribes certain proteins called as transcription factors which act as cis-acting element in enhancing the promoter activity thereby increasing the gene expression of the concerned gene.
Thus abiotic stress tolerant crops could be engineered by [1] successful fine-tuning of the stress response by engineering novel regulatory targets; [2] proper knowledge of posttranslational modifications which regulate plant growth performance under stress; [3] overexpression of miRNAs or their targets; [4] maintaining hormone homeostasis to avoid pleiotropic effects under stress; and [5] applying plant synthetic biology approaches to improve genetic engineering strategies (Abogadallah 2010); thus this could be achieved by altering the gene expression by altering the transcription factors, which is more or less like epigenetic modifications as there is no alteration of gene sequence. The tight regulation and fine tuning of TF genes during plant stress contribute to the foundation of complex signaling web and the important roles of NAC proteins are controlling gene expression through interacting with cis-elements, which are nothing but conserved sequence present upstream the promoter, which in turn is being induced by the respective TF involved as shown in example with rice [Figure 3] where the cis-acting elements( like NACR, DRE/CRT, ABRE, CE) and the signaling kinases that phosphorylate the concerned TFs and the carry out the gene expression of the Abiotic stress related genes.
Several recent reviews have addressed the role of specific TF families in abiotic stress responses which elucidates the possible mechanism to combat stress condition. In this review, the focus will be on selected TFs i.e., NAC from several large plant TF families, which have been shown to play significant roles in responses to the important abiotic stress factors, including drought, high salinity, high osmolarity, threshold temperatures and the chemical abiotic stress factor ABA.
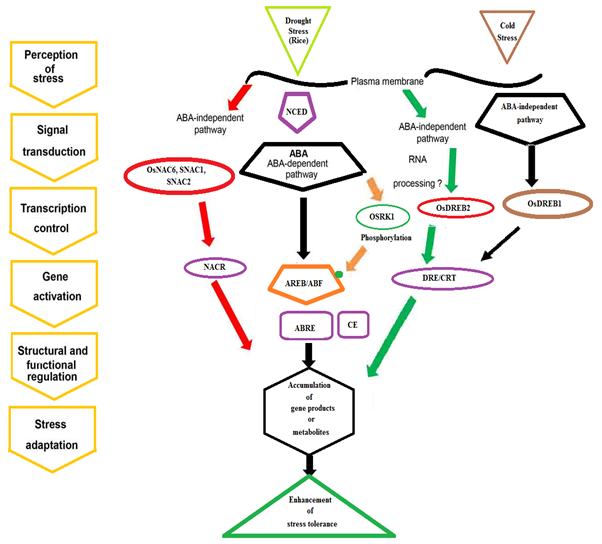
Figure 3: Transcriptional regulatory networks of the cis-elements NAC
transcription factors involved in abiotic stress-induced gene expression.
Adapted from Nuruzzaman., et al. 2013 PMC3759801
Engineering for transcription factors: Management of single gene level, for stress tolerance is not so affective; efforts were made to raise transgenic plants for stress inducible transcription factors: as a transcription factor regulates many genes. It is a possibility that many stress responsive genes, may share a common transcription factor. Various transcription factors belonging to the families of DREB, NAC, MYB, MYC, Cys2His2 zinc finger, bZIP, AP2/ERF and WRKY are known to be involved in abiotic stress tolerance. They bind to the promoter and/or regulatory elements of genes responsive to stress. Member (s) of different groups may be involved in a single response, and members of the same group may also be responsible for different kind of stress responses. Many transgenic tolerant to abiotic stresses have been produced through genetic engineering of gene (s) for transcription factors. The plant specific transcription factor group NAC (NAM, ATAF1/2, and CUC2) is required for its role in plant development and stress response. Transgenic rice plants overexpressing SNAC1 (stress responsive NAC1) showed enhanced salinity and drought tolerance (Hu., et al. 2006). Similarly, rice plants overexpressing SNAC2 (a rice NAC transcription factor group), exhibit higher salinity tolerance (Hu., et al. 2008). Overexpression of OsNAC5 in rice and Arabidopsis ectopic expression in enhanced salinity and drought tolerance, while knockdown of this gene in rice by RNAi lead to salt susceptibility (Song., et al. 2011). Likewise overexpression of ONAC045 encoding for NAC transcription factor gene in rice enhanced salinity and drought tolerance (Zheng., et al. 2009).
Genes responsible for Abiotic stress tolerance: Some of the transcription factors known to be responsible for abiotic stress tolerance and have been used for development of transgenic plants against various extreme environmental stresses are described below: There has been a continuous and tremendous research on transcriptional control of abiotic stress in model and crop plants. So to enumerate all that work is out of the scope of this review. More emphasis will be given on work done on drought, salinity and cold tolerance in regard to transcription factors especially family.
NAC (NAM ATAF and CUC) Transcription Factors: The NAC acronym is derived from three genes that were initially discovered to contain a particular domain (the NAC domain): NAM (for no apical meristem), ATAF1 and −2, and CUC2 (for cup-shaped cotyledon) (Souer., et al. 1996; Aida., et al. 1997). The NAC genes constitute one of the largest families of plant-specific TFs and are present in a wide range of species. Extensive investigation aided by the availability of several complete plant genomic sequences has identified 117 NAC genes in Arabidopsis, 151 in rice, 79 in grape, 26 in citrus, 163 in poplar, and 152 each in soybean and tobacco(Rushton., et al. 2008; Hu., et al. 2010; Nuruzzaman., et al. 2010, 2012a; Le., et al. 2011).
NAC Transcription Factors and their DNA Binding Structure: In Arabidopsis the DNA binding site of stress-inducible NAC TFs, the NAC recognition sequence (NACRS) containing the CACG core motif, has been identified by intensive analysis of the promoter of the drought inducible EARLY RESPONSE TO DEHYDRATION1 (ERD1) gene, which encodes a protein with homology to the ATP-binding subunit of the Clp ATP dependent protease from Escherichia coli. (Tran., et al. 2004; Simpson., et al. 2003). Molecular analysis of the ERD1 promoter demonstrated that ERD1 expression during dehydration mainly depends upon the integrity of both the “CAC TAA ATT GTC AC” ZFHDRS and the “ANN NNN TCN NNN NNN ACA CGC ATG T” NACRS sequences. (Tran., et al. 2004; Tran., et al. 2007), The CACG core motif is underlined. The rice drought-inducible ONAC TFs bind to a similar NACRS found in rice genome, demonstrating that the NACRS might be conserved across the plant kingdom at least for stress-inducible NAC TFs. (Hu., et al. 2006; Fang., et al. 2008). In addition, other sequences have also been reported as NAC binding sites. The Arabidopsis NAC1 protein has been shown to bind to a 21-bp segment (CTG ACG TAA GGG ATG ACG CAC) within the 35S -90 promoter fragment (Xie., et al. 2000). Independently; Duval., et al. 2002 demonstrated that purified AtNAM recombinant protein protected the region of the CaMV 35S promoter between -70 and -76, which is located in the 21-bp segment (AGGGATG). On the other hand, CBNAC TF of Arabidopsis bound specifically in vitro and in vivo to CBNACBS. Thus the NAC TFs contain a highly conserved N-terminal DNA-binding NAC domain and a variable transcriptional regulation C-terminal domain (Olsen., et al. 2005; Ernst., et al. 2004).
The presence of NAC domain in the NAC TFs was first revealed by database searches with the sequences of the cloned NAM, ATAF and CUC2 (Olsen., et al. 2005; Ooka., et al. 2004). Typically, the NAC domain is ~150 amino acids in length, and crystal structure of the NAC domain of an Arabidopsis NAC protein, the ANAC (ABA responsive NAC), was determined (Ernst., et al. 2004; Valliyodan., et al.).
NACs and their role in abiotic stress Response through Transgenic Development: NAC transcription factors play important roles in plant growth, development and stress responses (Hao., et al. 2011) The NAC gene family encodes one of the largest families of plant specific transcription factors and is absent in other eukaryotes (Gao., et al. 2007). Rice and Arabidopsis genomes contain 75 and 105 putative NAC genes, respectively for example, 33 NAC genes changed significantly in Arabdopsis under salt treatment (Jiang and Deyholos, 2006), 38 NAC genes were involved in response to drought in soybean (Le., et al. 2011), 40 NAC genes responded to drought or salt stress in rice (Fang., et al. 2008), 32 NAC genes responded to at least two kinds of treatments in Chrysanthemum lavandulifolium (Huang., et al. 2012). It appears that a significant proportion of NAC genes function in stress response according to the expression data from genome-wide transcriptome analyses in many plants. The role of NAC gene family in abiotic stresses was discovered in Arabidopsis.
Three NAC genes were induced under salt and/or drought stress. Overexpression of three genes ANACO19, ANACO55, ANACO72, greatly enhanced drought tolerance in Arabidopsis (Tran., et al. 2004). Multiple rice transcription factors, including a NAC gene were induced in the early stages of salt stress (Chao., et al. 2005) OsNAC6, a member of ATAF family, was also induced by cold, salt, drought and abscisic acid (ABA) (Ohnishi., et al. 2005). Overexpression of stress responsive gene SNAC1 (stress-responsive NAC1) significantly enhances drought resistance in transgenic rice (22-34% higher seed setting than control) in the field under severe drought stress conditions at the reproductive stage while showing no phenotypic changes or yield penalty. The other NAC Family SNAC3-OE transgenic plants exhibited significantly lower levels of H2O2, malondiadehyde (MDA), and relative electrolyte leakage than the wild-type control under heat stress conditions, implying that SNAC3 may confer stress tolerance by modulating reactive oxygen species (ROS) homeostasis. Quantitative PCR experiments showed that the expression of a large number of ROS-scavenging genes was dramatically increased in the SNAC3-OE plants, but significantly decreased in the SNAC3-RNAi transgenic plants (Xiong., et al. 2015).
The transgenic rice also shows significantly improved drought resistance and salt tolerance at the vegetative stage. Plant TFs of different families, e.g., NAC, bHLH, MYB, bZIP, WRKY, ZF, and AP2/ERF also have significant degrees of ID, which can play a functional role in interactions with other regulatory proteins, as exemplified by the interaction between the C-terminal domain of HvNAC013 and Radical Induced Cell Death1 (RCD1) (Kjaersgaard., et al. 2011; Kragelund., et al. 2012). Compared with wild type, the transgenic rice were more sensitive to abscisic acid and lost water more slowly by closing more stomatal pores, yet displayed no significant difference in the rate of photosynthesis. DNA chip analysis revealed that a large number of stress related genes were up-regulated in the SNAC1-overexpressing rice plants. SNAC1 holds promising utility in improving drought and salinity tolerance in rice (Hu., et al. 2006).
Similarly several NAC genes are found to be induced by drought, high salinity, and cold stresses. OsNAC6/SNAC2 encodes a NAC transcription factor in rice (Hu., et al. 2008). Expression of OsNAC6/SNAC2 is induced by cold, drought and high salinity. Transgenic rice plants overexpressing OsNAC6 showed enhanced tolerance to dehydration and high salinity stresses, although the plants showed growth retardation and low yield under the non-stress condition. This low yield was increased in transgenic rice plants driven by a stress-inducible promoter. These results indicate OsNAC6 functions as a transcriptional activator in abiotic stress responses (Chung., et al. 2009). In a similar experiment OsNAC5 expression was induced by dehydration, cold, ABA, and MeJA treatments. The growth of rice plants overexpressing OsNAC5 was comparable to that of control plants under non-stress conditions. The OsNAC5 over expressers exhibited enhanced tolerance to high salinity. In another study it was reported that RNA interference (RNAi) transgenic rice plants with reduced OsNAC5 expression exhibited a reduced amount of tolerance to abiotic stresses than control plants, whereas overexpression of OsNAC5 improved abiotic stress tolerance. It was also revealed that accumulation of compatible solutes such as proline and soluble sugars was absolutely linked with OsNAC5 expression level (Song., et al. 2011). A recent report revealed that GmNAC11 acts as a transcriptional activator, whereas GmNAC20 functions as a mild repressor; however, the C-terminal end of GmANC20 has transcriptional activation activity.
Over-expression of GmNAC20 increased salt and freezing tolerance in transgenic Arabidopsis plants; however, over-expression only improved salt tolerance. Over-expression of GmNAC20also enhanced lateral root formation (Hao., et al, 2011). Thus the above study revealed that GmNAC20 may regulate stress tolerance through activation of the DREB/CBF–COR pathway, and may control lateral root development by altering auxin signalingrelated genes. Recently, an attempt was made to introduce NAC1 gene from E. coli into tobacco through Agrobacteriummediated transformation (Ramegowda., et al. 2012). In another experiment, overexpression of TaNAC2 resulted in enhanced tolerances to drought, salt, and freezing stresses in Arabidopsis, which were simultaneously demonstrated by enhanced expression of abiotic stress-response genes and several physiological indices (Mao., et al. 2012). The transgenic plants resulted in increased tolerance towards abiotic stresses.
The above results thus indicate that NAC genes also play an important role in the transcriptional complex of abiotic stress responses in plants not only this NAC also plays important role in senescence, Overexpression of AtNAP[58], ORE1 (Kim., et al. 2009; Balazadeh., et al. 2010), ORS1 (Balazadeh., et al. 2011), ANAC016 (Kim., et al. 2013), and ATAF1 (Garapati., et al. 2015) resulted in precocious senescence, whereas blocking the function of these genes delayed senescence suggesting that they function as nonredundant positive regulators of senescence in Arabidopsis so NAC transcription factors are extensively implicated in the senescence process as judged from global gene expression analysis which show their diversity of function (Skriver., et al. 2015). So understanding the molecular mechanisms of NAC TFs networks integrating multiple stress responses will be essential for the development of broad-spectrum stress tolerant crop plants that can better cope with environmental challenges in future climates (Shao., et al. 2015).
| Gene | Species | Targeted Trait | References |
| AhNAC2 | Arachis pintoi | Salinity Stress | Liu., et al. 2011 |
| ATAF1 | Arabidopsis thaliana | Drought Stress | Lu., et al. 2007 |
| AtNAC2 | Arabidopsis thaliana | Salinity Stress | He., et al. 2005 |
| AtNAC019 | Arabidopsis thaliana | Drought and Salt Stress | Lu., et al. 2007 |
| AtNAC055 | Arabidopsis thaliana | Drought and Salt Stress | He., et al. 2005 |
| AtNAC072 | Arabidopsis thaliana | Drought and Salt Stress | Tran., et al. 2004 |
| ANAC013 | Arabidopsis thaliana | Drought and Salt Stress | Clercq., et al. 2013 |
| ONAC063 | Arabidopsis thaliana | High Salinity and Osmotic Stress | Yokotani., et al. 2009 |
| TaNAC2 | Arabidopsis thaliana | Drought and Salt Stress | Mao., et al. 2012 |
| TaNAC29 | Arabidopsis thaliana | Salt, drought, Senescence Process | Huang., et al. 2015 |
| NTL6, SnRK2.8 | Arabidopsis thaliana | Drought Stress | Kim., et al. 2012 |
| CsNAM | Camellia sinensis | Drought, Osmoticum, Salt and Heat Stress | Paul., et al. 2012 |
| CarNAC5 | Cicer arietinum | Drought and Salt Stress | Yu., et al. 2015 |
| CarNAC5 | Cicer arietinum | Drought and Heat Stress | Peng., et al. 2009 |
| GhNAC3 | Gossypium hirsutum | Cold Stress | Meng., et al. 2009 |
| GhNAC4 | Gossypium hirsutum | Drought Cold and Salt Stress | Meng., et al. 2009 |
| GhNAC5 | Gossypium hirsutum | Cold and Drought Stress | Meng., et al. 2009 |
| GhNAC6 | Gossypium hirsutum | Cold, Drought and Salt Stress | Meng., et al. 2009 |
| SNAC1 | Gossypium hirsutum | Drought and salinity Stress | Liu., et al. 2014 |
| GmNAC11, GmNAC20 | Glycine max | Drought and salinity Stress | Hao., et al. 2011 |
| GmNAC11 | Glycine max | Salinity Stress | Hao., et al. 2011 |
| HvNAC6 | Hordeum vulgare | ABA mediated defense responses | Chen., et al. 2013 |
| TaNAC2a | Nicotiana tabacum | Drought Stress | Tang., et al. 2012 |
| OsNAC5/ Os11g08210 | Oryza sativa | ABA, salt, cold tolerance, grain filling Stress | Sperotto., et al. 2009 |
| OsNAC9 | Oryza sativa | Drought Stress | Kim., et al. 2012 |
| OsNAC10/Os11g03300 | Oryza sativa | Root, panicle, drought, salt, ABA | Jeong., et al. 2010 |
| OsOAT, SNAC2 | Oryza sativa | Drought and oxidative stress tolerance | You., et al. 2013 |
| OsEREBP1 | Oryza sativa | Abiotic and biotic stress | Jisha., et al. 2015 |
| SNAC1 | Oryza sativa | Cold, Drought, and Salt Stress | Hu., et al. 2006 |
| SNAC3 | Oryza sativa | Drought and Salt Stress | Fang., et al. 2015 |
| OsNAC6 | Oryza sativa | Drought, Salt Stress | Nakahsima., et al. 2007 |
| OsNAP | Oryza sativa | Salinity, drought and low temperature Stress | Chen., et al. 2014 |
| ATNAC3 | Solanum lycopersicum | Drought and Salt Stress | Abdallat., et al. 2015 |
| SlNAC4 | Solanum Lycopersicum | Drought and Salt Stress | Zhu., et al.2014 |
| SiNAC | Setaria italica | Drought and Salt Stress | Puranik., et al.2011 |
| TaNAC4 | Triticum aestivum | Cold, Salt, and Wounding Stress | Xia., et al.2010 |
| SNAC1 | Triticum aestivum | Drought and salinity Stress | Yu., et al.2013 |
| TaNAC4a, TaNAC6 | Triticum aestivum | Dehydration, salinity and low temperature Stress | Tang., et al.2012 |
| ZmSNAC1 | Zea. mays | Low temperature, Drought and Salinity Stress | Lu., et al.2012 |
Table 1: Transgenic plants developed for Abiotic stress tolerance by over-expression of NAC genes.
Conclusion
We also are aware of the fact that this stress tolerance cannot be achieved only by single gene as the abiotic stress is a polygenic trait, so for constitutive expression of these genes can be achieved by strong constitutive promoter but it has its own problems of energy loss and deleterious effects but if we can control gene expression a well regulated expression of stress-induced transcription factors, which would regulate the expression of a large no of concerned downstream genes. Considerable information through transcriptome profiling strategies has been gained about NAC TFs since the discovery of them, but the research in this area is still in its initial phase. Genome-wide identification and expression profiling has undoubtedly open new avenues for describing the key features of TFs. As a result, our current understandings of the regulatory functions of the TFs in various plant species will be definitely accelerated. In particular, the stress responsive TFs can be used as promising candidates for generation of stress tolerant transgenic plants possessing high productivity under adverse conditions. But there are still some problems to be solved as some studies have shown. Firstly, the constitutive overexpression of TF genes occasionally may lead to negative effects in transgenic plants such as dwarfing, late flowering and lower yields. Secondly, the transgenic plants overexpressing genes may occasionally have antagonistic responses to different stresses. Thirdly, very few of transgenic plants overexpressing NAC TFs genes were evaluated in the field trials so far.
Thus as everyone knows, the prime aims for plant stress research is to provide targets for the improvement of stress tolerance in crop plants. With the unforeseen changes in climatic conditions will lead to more complex stress environment in the fields, we will face new hurdles in creating the multiple stress-tolerant crops. Targeting transcription factors are safe as it is just like epigenetic switching on or off of the gene expressing machinery. Future research should focus on additional approaches including: 1) better knowledge of the complex network of genes and the resultant metabolic and physiological changes during stress tolerance; 2) Better understanding of the molecular determinants that simultaneously regulate responses to stress and plant developmental processes; 3) combining different approaches with multiple gene engineering to target diverse stress response mechanisms and obtain the desired traits and effects; 4) optimizing TF technologies through reducing growth abnormalities and enhancing the introduced trait to withstand stress, thus taking advantage of available high throughput tools such as transcriptomics, proteomics and Metabolomics. Finally, the collective and cooperative efforts of plant molecular biologists, physiologists, and breeders are required to generate stress-tolerant grasses through genetic engineering. It is hoped that, in the future, these collective efforts and results of collaborative studies will positively contribute to sustainable food production in the world and will help to prevent global-scale environmental damage that results from abiotic stress.
Acknowledgment
The authors express their acknowledgements to the Woman Scientist scheme-A, Department of Science and Technology, Ministry of Science and Technology, and SVP University of Agriculture and Technology Meerut India for the financial and technical support to conduct activities on research program.
The authors express their acknowledgements to the Woman Scientist scheme-A, Department of Science and Technology, Ministry of Science and Technology, and SVP University of Agriculture and Technology Meerut India for the financial and technical support to conduct activities on research program.
References
- Abdallat AMA., et al. “Overexpression of the Transcription Factor HvSNAC1 Improves Drought Tolerance in Barley (Hordeum vulgare L.)”. Molecular Breeding33.2 (2014): 401-414.
- Abogadallah GM. “Antioxidative defense under salt stress”. Plant Signaling and Behavior5.4 (2010): 369-374.
- Agarwal M., et al. “A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance”. Journal of Biological Chemistry 281.49 (2006): 37636-37645.
- Agarwal PK., et al. “Role of DREB transcription factors in abiotic and biotic stress tolerance in plants”. Plant Cell Reports25.12 (2006): 1263-1274.
- Agarwal P., et al. “Overexpression of PgDREB2A transcription factor enhances abiotic stress tolerance and activates downstream stress-responsive genes”. Molecular Biology Reports 37.2 (2010): 1125-1135.
- Aida M., et al. “Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant”. Plant Cell 9.6 (1997): 841-857.
- Balazadeh S., et al. “ORS1, an H2O2-Responsive NAC Transcription Factor, Controls Senescence in Arabidopsis thaliana”. Molecular Plant 4.2 (2011): 346-360.
- Balazadeh S., et al. “Mueller-Roeber, B. A gene regulatory network controlled by the NAC transcription factor ANAC092/AtNAC2/ORE1 during salt-promoted senescence”. The Plant Journal 62.2 (2010): 250-264.
- Cabello JV., et al. “Novel perspectives for the engineering of abiotic stress tolerance in plants”. Current Opinion in Biotechnology 26 (2014): 62-70.
- Chao DY., et al. “Salt-responsive genes in rice revealed by cDNA microarray analysis”. Cell Research 15.10 (2005): 796-810.
- Chung PJ., et al. “The histone deacetylase OsHDAC1 epigenetically regulates the OsNAC6 gene that controls seedling root growth in rice”. The Plant Journal 59.5 (2009): 764-776.
- Fang Y., et al. “Systematic sequence analysis and identification of tissue-specific or stress-responsive genes of NAC transcription factor family in rice”. Molecular Genetics and Genomics280.6 (2008): 547-563.
- Feist AM and Palsson BO. “The growing scope of applications of genome-scale metabolic reconstructions using Escherichia coli”. Nature Biotechnology 26.6 (2008): 659-667.
- Gao J P., et al. “Understanding abiotic stress tolerance mechanisms: recent studies on stress response in rice”. Journal of Integrative Plant Biology 49.6 (2007): 742-750.
- Garapati P., et al. “Transcription Factor ATAF1 in Arabidopsis Promotes Senescence by Direct Regulation of Key Chloroplast Maintenance and Senescence Transcriptional Cascades”. Plant Physiology 168.3 (2015):1122-1139.
- Golldack D., et al. “Plant tolerance to draught and salinity: Stress regulating transcription factors and their functional significance in the cellular transcriptional network”. Plant Cell Reports 30.8 (2011): 1389-1391.
- Han X., et al. “Two NAC transcription factors from Caragana intermedia altered salt tolerance of the transgenic Arabidopsis”. BMC Plant Biology 15 (2015): 208.
- Hu H., et al. “Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice”. Plant Molecular Biology 67.2 (2008): 169-181.
- Hu R., et al. “Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa”. BMC Plant Biology 10 (2010): 145.
- Huang H., et al. “Transcriptome-wide survey and expression analysis of stress-responsive NAC genes in Chrysanthemum lavandulifolium”. Plant Science 193 (2012): 18-27.
- Inge De Clercq., et al. “The Membrane-Bound NAC Transcription Factor ANAC013 Functions in Mitochondrial Retrograde Regulation of the Oxidative Stress Response in Arabidopsis”. The Plant Cell 25.9 (2013): 3472-3490.
- Jiang Y., et al. “Comprehensive transcriptional profiling of NaCl-stressed Arabidopsis roots reveals novel classes of responsive genes”. BMC Plant Biology 6 (2006): 25.
- Katerji N., et al. “Salinity effect on crop development and yield, analysis of salt tolerance according to several classification methods”. Agricultural Water Management 62.1 (2003): 37-66.
- Katerji N., et al. “Comparison of corn yield response to plant water stress caused by salinity and drought”. Agricultural Water Management 65.2 (2004): 95-101.
- Kim YS and Kim JK. “Rice transcription factor transcription factor AP37 involved in grain yield increase under drought stress”. Plant Signaling and Behavior 4.8 (2009): 735-736.
- Kim YS., et al. “Mutation of the Arabidopsis NAC016 transcription factor delays leaf senescence”. Plant and Cell Physiology 54.10 (2013): 1660-1672.
- Kjaersgaard T., et al. “Senescence-associated barley NAC (NAM, ATAF1,2, CUC) transcription factor interacts with radical-induced cell death 1 through a disordered regulatory domain”. The Journal of Biological Chemistry286.41 (2011): 35418-35429.
- Kragelund BB., et al. “Order by disorder in plant signaling”. Trends in Plant Science17.11 (2012): 625-632.
- Le DT., et al. “Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress”. DNA Research 18.4 (2011): 263-276.
- M Ashraf. “Biotechnological approach of improving plant salt tolerance using antioxidants as markers”. Biotechnology Advances 27.1 (2009): 84-93.
- Magnani E., et al. “From endonuclease to transcription factors: Evolution of the AP2 DNA binding domain in plants”. Plant Cell16.9 (2004): 2265-2277.
- Mao Xinguo., et al. “Novel NAC Transcription Factor TaNAC67 Confers Enhanced Multi-Abiotic Stress Tolerances in Arabidopsis”. PLoS ONE 9.1 (2014): e84359.
- Nambara E., et al. “Arabidopsis transcriptome analysis under drought, cold, high-salinity and ABA treatment conditions using a tiling array”. Plant and Cell Physiology 49.8 (2008): 1135-1149.
- Nuruzzaman M., et al. “Genome-wide analysis of NAC transcription factor family in rice”. Gene 465.1 (2010): 30-44.
- Nuruzzaman M., et al. “A genome-wide survey of the conditions”. Plant Signaling & Behavior 7 (2010): 1518-1521.
- Oh SJ., et al. “Expression of barley HvCBF4 enhances tolerance to abiotic stress in transgenic rice”. Plant Biotech Journal 5.5 (2007): 646-656.
- Ohnishi T., et al. “OsNAC6, a member of the NAC gene family, is induced by various stresses in rice”. Genes & Genetic Systems 80.2 (2005): 135-139.
- Olsen AN., et al. “NAC transcription factors: structurally distinct, functionally diverse”. Trends in Plant Science 10.2 (2005): 79-87.
- Qin F., et al. “Achievements and challenges in understanding plant abiotic stress responses and tolerance”. Plant and Cell Physiology 52.9 (2011): 1569-1582.
- Riechmann JL., et al. “Arabidopsis transcription factors: genome –wide comparative analysis among eukaryotes”. Science 290.5499 (2000): 2105-2110.
- Riechmann JL., et al. “The AP2/EREBP family of plant transcription factors”. Biological Chemistry379.6 (1998): 633-646.
- Rushton PJ., et al. “Tobacco transcription factors: novel insights into transcriptional regulation in the Solanaceae”. Plant Physiology 147.1 (2008): 280-295.
- SC Saxena., et al. “Osmoprotectants: Potential for Crop Improvement Under Adverse Conditions.” Plant Acclimation to Environmental Stress(2012): 197-232.
- Shao H., et al. “NAC transcription factors in plant multiple abiotic stress responses: progress and prospects”. Frontiers in Plant Science 6 (2015): 902.
- Simpson SD., et al. “Two different novel cis-acting elements of erd1, a clpA homologous Arabidopsis gene, function in induction by dehydration stress and dark-induced senescence”. The Plant Journal33.2 (2003): 259-270.
- Skriver K., et al. “NAC Transcription Factors in Senescence: From Molecular Structure to Function in Crops.” Plants 4.3 (2015): 412-448.
- Souer E., et al. “The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries”. Cell 85.2 (1996): 159-170.
- Sakuma Y., et al. “DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression”. Biochemical and Biophysical Research Communications 290.3 (2002): 998-1009.
- Valliyodan B., et al. “Understanding regulatory networks and engineering for enhanced drought tolerance in plants”. Current Opinion in Plant Biology 9.2 (2006): 189-195.
- Xiong L., et al. “A stress-responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice”. Journal of Experimental Botany 66.21 (2015): 6803-6817.
- Xiong L., et al. “Regulation of osmotic stress responsive gene expression by LOS6/ABA1 locus in Arabidopsis”. The Journal of Biological Chemistry 277.10 (2020): 8588-8596.
- Yamaguchi-Shinozaki K and K Shinozaki. “A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress”. Plant Cell 6.2 (1994): 251-264.
- Yujie Fang., et al. “A stress-responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice”. Journal of Experimental Botany 66.21 (2015): 6803-6817.
- Zhang JZ., et al. “From laboratory to field: Using information from Arabidopsis to engineer salt, cold, and drought tolerance in crops”. Plant Physiology 135.2 (2004): 615-621.
- Zhang X., et al. “OsDREB2A, a rice transcription factor, significantly affects salt tolerance in transgenic soybean”. PLoS ONE 8.12 (2013): 83011-83020.
- Zhang Y., et al. “Expression of a rice DREB1 gene, OsDREB1D, enhances cold and high salt tolerance in transgenic Arabidopsis”. Biochemistry and Molecular Biology Reports 42.8 (2009): 486-492.
- Liu G., et al. “Overexpression of rice NAC gene SNAC1 improves drought and salt tolerance by enhancing root development and reducing transpiration rate in transgenic cotton”. PLoS ONE 9.1 (2014): e86895.
- Liu L., et al. “Cloning of a Vacuolar H(+)-pyrophosphatase Gene from the Halophyte Suaeda corniculata whose Heterologous Overexpression Improves Salt, Saline-alkali and Drought Tolerance in Arabidopsis”. Journal of Integrative Plant Biology 53.9 (2011): 731-742.
- Lu PL., et al. “A novel drought-inducible gene, ATAF1, encodes a NAC family protein that negatively regulates the expression of stress-responsive genes in Arabidopsis”. Plant Molecular Biology 63.2 (2007): 289-305.
- He XJ., et al. “AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development”. The Plant Journal 44.6 (2005): 903-916.
- Huang Q., et al. “TaNAC29, a NAC transcription factor from wheat, enhances salt and drought tolerance in transgenic Arabidopsis”. BMC Plant Biology 15 (2015): 268.
- Hao YJ., et al. “Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants”. The Plant Journal 68.2 (2011): 302-313.
- Hu HH., et al. “Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice”. Proceedings of the National Academy of Sciences USA 103.35 (2006): 12987-12992.
- Tran LSP., et al. “Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter”. Plant Cell 16.9 (2004): 2481-2498.
- Mao X., et al. “TaNAC2, a NAC-type wheat transcription factor conferring enhanced multiple abiotic stress tolerances in Arabidopsis”. Journal of Experimental Botany 63.8 (2012): 2933-2946.
- Meng C., et al. “Characterization of six novel NAC genes and their responses to abiotic stresses in Gossypium hirsutum L”. Plant Science 176.3 (2009): 352-359.
- Kim JK., et al. “The overexpression of OsNAC9 alters the root architecture of rice plants enhancing drought resistance and grain yield under field conditions”. Plant Biotechnology Journal 10.7 (2012): 792-805.
- Peng H., et al. “Characterization of a chickpea (Cicer arietinum L.) NAC family gene, CarNAC5, which is both developmentally- and stress-regulated”. Plant Physiology & Biochemistry 47.11 (2009): 1037-1045.
- Puranik S., et al. “Molecular cloning and characterization of a membrane associated NAC family gene, SiNAC from foxtail millet [Setaria italica (L.) P. Beauv.]”. Molecular Biotechnology 49.2 (2011): 138-150.
- Yu X., et al. “A chickpea stressresponsive NAC transcription factor, CarNAC5, confers enhanced tolerance to drought stress in transgenic Arabidopsis”. Plant Growth Regulation 79.2 (2016): 187-197.
- Chen X., et al. “The NAC family transcription factor OsNAP confers abiotic stress response through the ABA pathway”. Plant and Cell Physiology 55.3 (2004): 604-619.
- Fang W., et al. “GmWRKY27 interacts with GmMYB174 to reduce expression of GmNAC29 for stress tolerance in soybean plants”. The Plant Journal 83.2 (2015): 224-236.
- Abdallat AMA., et al. “Overexpression of two ATNAC3 related genes improves drought and salt tolerance in tomato (Solanum lycopersicum L.)”. Plant Cell, Tissue and Organ Culture 120.3 (2015): 989-1001.
- Zhu M., et al. “The Abiotic stress responsive NAC Type Transcription Factor SlNAC 4 regulates salt and drought tolerance and stress related genes in tomato S. Lycopersicum”. Plant Cell Reports 33.11 (2014): 1851-1863.
- Xia N., et al. “Characterization of a novel wheat NAC transcription factor gene involved in defense response against stripe rust pathogen infection and abiotic stresses”. Molecular Biology Reports 37.8 (2010): 3703-3712.
- Yu Cai L., et al. “A rice stress-responsive NAC gene enhances tolerance of transgenic wheat to drought and salt stresses”. Plant Science 203 (2013): 33-40.
- Abogadallah GM. “Antioxidative defense under salt stress”. Plant Signaling and Behavior 5.4 (2010): 369-374.
- Manavalan LP., et al. “Physiological and molecular approaches to improve drought resistance in soybean”. Plant and Cell Physiology 50.7 (2009): 1260-1276.
- Udvardi MK., et al. “Legume transcription factors: Global regulators of plant development and response to the environment”. Plant Physiology 144.2 (2007): 538-549.
- Umezawa T., et al. “Engineering drought tolerance in plants: discovering and tailoring genes to unlock the future”. Current Opinion in Plant Biotechnology 17.2 (2006): 113-122.
- Yadav S., et al. “Salinity Tolerance in plants”. Biotech Today 3.2 (2013): 53-56.
- Flowers TJ and Yeo AR. “Breeding for salinity resistance in crop plants-where next?” Australian Journal of Plant Physiology 22 (1995): 876-884.
- Fowler S and Thomashow MF. “Arabidopsis transcriptome profiling indicates that multiple regulatory pathways are activated during cold acclimation in addition to the CBF cold response pathway”. The Plant Cell 14.8 (2002): 1675-1690.
- Katerji N., et al. “Salinity effect on crop development and yield, analysis of salt tolerance according to several classification methods”. Agricultural Water Management 62.1 (2003): 37-66.
- Katerji N., et al. “Comparison of corn yield response to plant water stress caused by salinity and drought”. Agricultural Water Management 65.2 (2004): 95-101.
- Chaves M.M., et al. “Recent Advances in Photosynthesis Under Drought and Salinity”. Advances in Botanical Research 57 (2011): 49-104.
- Basu U. “Identification of Molecular Processes Underlying Abiotic Stress Plants Adaptation Using 'Omics' Technologies”. Sustainable Agriculture and New Biotechnologies (2012): 149-172.
- Grover A., et al. “Understanding molecular alphabets of the plant abiotic stress responses”. Current Science 80.2 (2001): 206-216.
- Matsui A., et al. “Arabidopsis transcriptome analysis under drought, cold, high-salinity and ABA treatment conditions using a tiling array”.Plant and Cell Physiology 49.8 (2008): 1135-1149.
- Cutler SR, et al. “Abscisic acid: emergence of a core signaling network”. Annual Reviews in Plant Biology 61 (2010): 651-679.
- Finkelstein., et al. “Abscisic Acid Signaling in Seeds and Seedlings”. Plant Cell 14 (2002): S15-S45.
- Kavar T., et al. “Identification of genes involved in the response of leaves of Phaseolus vulgaris to drought stress”. Molecular Breeding 21 (2007): 159-172.
- Bohnert HJ., et al. A genomics approach towards salt stress tolerance”. Plant Physiology and Biochemistry 39 (2001): 295-311.
- E Nomura M., et al. “A genomic approach towards salt stress tolerance”. Plant Physiology and Biochemistry 39.3 (2001): 295-311.
- Seki M., et al. “Transcriptome analysis in abiotic stress conditions in higher plants”. Topics in Current Genetics 4 (2003): 271-295.
- Seki M., et al. “Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray”. The Plant Journal 31.3 (2002): 279-292.
- Shinozaki K and Yamaguchi-Shinozaki K. “Gene networks involved in drought stress tolerance and response”.Journal of Experimental Botany 58.2 (2007): 221-227.
- Kilian J., et al. “Prerequisites, performance and profits of transcriptional profiling the abiotic stress response”. Biochimica et Biophysica Acta 1819.2 (2012): 166-175.
- Ooka H., et al. “Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana”. DNA Research10.6 (2003): 239-247.
- Xie Q., et al. “Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development”. Genes & Development14.23 (2000): 3024-3036.
- Ernst HA., et al. “Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors”. EMBO Reports5.3 (2004): 297-303.
- Duval M., et al. “Molecular characterization of AtNAM: A member of the Arabidopsis NAC domain superfamily”. Plant Molecular Biology 50.2 (2002): 237-248.
- Tran LSP., et al. “Co-expression of the stressinducible zinc finger homeodomain ZFHD1 and NAC transcription factors enhances expression of the ERD1 gene in Arabidopsis”. The Plant Journal 49.1 (2007): 46-63.
- Hu H., et al. “Overexpressing a NAM, ATAF and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice”. Proceedings of the National Academy of Sciences USA 103.35 (2006): 12987-12992.
- Song SY., et al. “Physiological mechanisms underlying OsNAC5-dependent tolerance of rice plants to abiotic stress”. Planta234.2 (2011): 331-345.
- Zheng X., et al. “Overexpression of a NAC transcription factor enhances rice drought and salt tolerance”. Biochemical and Biophysical Research Communications 379.4 (2009): 985-989.
- Yokotani N., et al. “Expression of rice heat stress transcription factor OsHsfA2e enhances tolerance to environmental stresses in transgenic Arabidopsis”. Planta 227.5 (2008): 957-967.
- Paul A., et al. “CsNAM-like protein encodes a nuclear localized protein and responds to varied cues in tea [Camellia sinensis (L.) O. Kuntze]”. Gene502.1 (2012): 69-74.
- Chen YJ., et al. “The barley HvNAC6 transcription factor affects ABA accumulation and promotes basal resistance against powdery mildew”. Plant Molecular Biology 83.6 (2013): 577-590.
- Sperotto RA., et al. “Identification of up-regulated genes in flag leaves during rice grain filling and characterization ofOsNAC5, a new ABA-dependent transcription factor”. Planta 230 (2009): 985-1002.
- Jeong JS., et al. “Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions”. Plant Physiology 153.1 (2010): 185-197.
- You J., et al. “The SNAC1-targeted gene OsSRO1c modulates stomatal closure and oxidative stress tolerance by regulating hydrogen peroxide in rice”. Journal of Experimental Botany 64.2 (2013): 569-583.
- Jisha V., et al. “Overexpression of an AP2/ERF Type Transcription Factor OsEREBP1 Confers Biotic and Abiotic Stress Tolerance in Rice”. PLoS One 10.6 (2015 ): e0127831.
- Nakashima K., et al. “Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice”. The Plant Journal 51.4 (2007): 617-630.
- Tang Y., et al. “Molecular characterization of novel TaNAC genes in wheat and overexpression of TaNAC2a confers drought tolerance in tobacco”. Physiologia Plantarum 144.3 (2012): 210-224.
- Lu M., et al. “A maize stress-responsive NAC transcription factor, ZmSNAC1, confers enhanced tolerance to dehydration in transgenic Arabidopsis”. Plant Cell Reports 31.9 (2012): 1701-1711.
Citation:
Ashu Singh., et al. “Abiotic Stress Tolerance in Crop Plants: With Respect to NAC Transcription Factor”. Innovative Techniques
in Agriculture 1.1 (2017): 63-76.
Copyright: © 2017 Ashu Singh., et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.





























 Scientia Ricerca is licensed and content of this site is available under a Creative Commons Attribution 4.0 International License.
Scientia Ricerca is licensed and content of this site is available under a Creative Commons Attribution 4.0 International License.